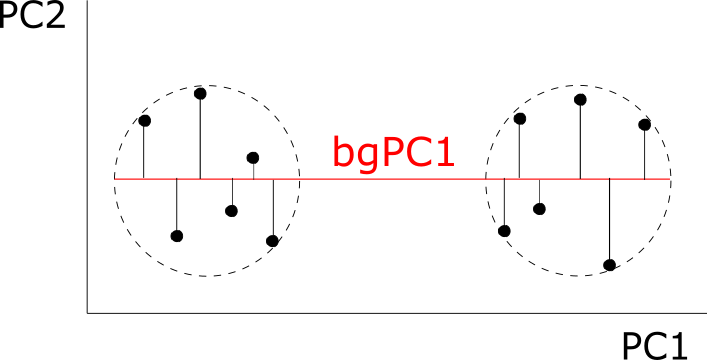
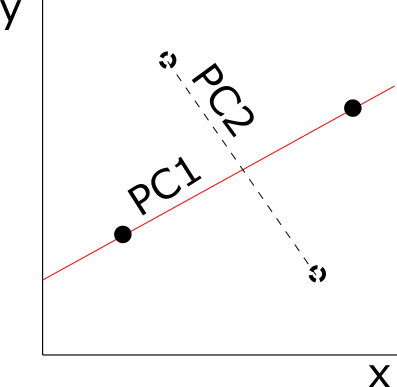
I once read a paper where the author said that shape’s a multidimensional and multivariate character. That let me thinking for months on the difference between these two terms. The next summer, during an oral presentation, a different scientist repeated that same sentence, this time explaining that shape’s a multidimensional character due to the number of variables that are used to describe it and a multivariate character due to the statistical treatment this character must receive to be analysed. Contrary to univariate analyses, multivariate techniques take into account multiple variables at the same time, meaning that they consider the correlation among variables (although they often assume the independence among them). Once we have many variables describing a phenotype, we can either apply one multivariate method to analyze this phenotype all at once or we can apply multiple univariate methods to decompose this phenotype in unrelated chunks of data.
As far as I’ve read, over the last 60 years there was a controversy in the scientific literature about which approach was optimal on the analyses of multidimensional data. Today, the vast majority of scientists (at least in geometric morphometrics) consider multivariate methods the standard, the most powerful approach to analyze shape data. However, not so long ago I read a paper that stroke me for its simplicity and how well it explained the difficult relationship between multivariate and univariate approaches: Healy (1969) (I hope you didn’t expect a recent paper).
I liked it so much that I replicated the figure showing the whole argument:
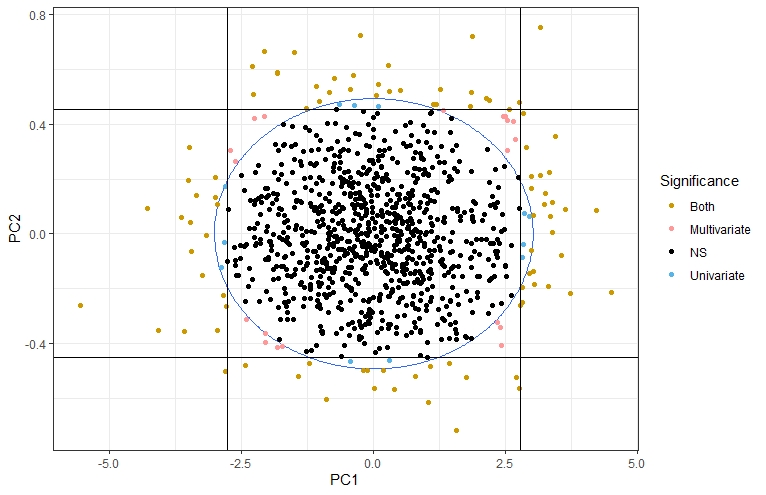
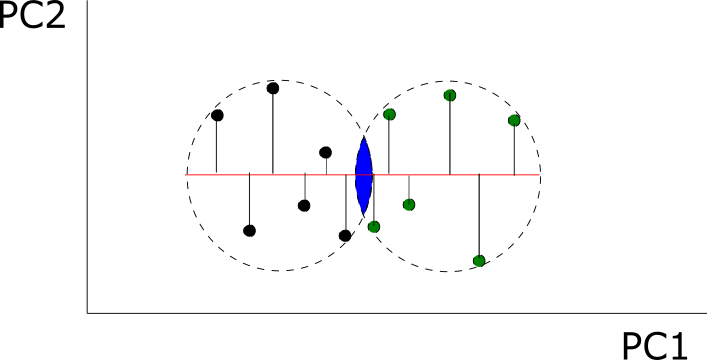
Take 1000 random observations from a bivariate normal distribution where each variable has variance 1 and covariance 0.95. Then, run a PCA for the whole sample and for each observation run a univariate test for each PC and a multivariate test for both variables at the same time. You get four different situations (represented in the figure): a) the observation isn’t significantly different from the rest of the population for any test (in black), b) the observation is significantly different only in its PC1 (in blue, observations off the vertical lines) or PC2 (in blue, observations off the horizontal lines), c) the observation is significantly different only in the multivariate test (in pink, off the blue circle but within the limits of the vertical and horizontal lines), d) significantly different for all tests (in gold, off the circle and the vertical and horizontal lines).
I learned a lot with these simple simulations: I could increase the number of variables, the patterns of covariation, etc. The most valuable lesson I got was that multivariate techniques don’t recover all the statistical signals identified by the univariate methods: multivariate methods are more sensitive in the direction of the covariance (the corners in the figure), while univariate methods detect more often observations in the direction of each axis (aligned with the PCs, as you can see). These conclusions may not be relevant for shape data, where all coordinates are a function of the whole configuration of coordinates (due to the Procrustes superimposition) and that would invalidate theoretically its decomposition in multiple chunks of independent data. Still, an entertaining learning experience.
Edit (4/10/20):
I attach here the original article by Rao, the one nicely explained by Healy (1969) and that are usually cited together. I’m posting it here since I really struggled to find it: I had to visit several libraries in Paris in very particular conditions, which made a funny memory from my postdoc there. Psychologists say that memories are created more easily when the learning experience is coupled with a physical one, maybe that’s why I like this whole thing. Anyway, here you’ve got it:
(It’s a large file, so it may take few seconds to download)